I just finished reading an excellent book on meta-analysis, Introduction to Meta Analysis by Borenstein et al.
The book starts with the motivating example of streptokinase, a blood clot buster. Early studies showed conflicting evidence of efficacy of the drug in preventing death following a heart attack. These early studies were small, and the main outcome was often not significantly different from zero; it took very large scale studies in 1986 and 1987 to confirm the protective effect of streptokinase, which could reduce deaths by 20%.
The book makes the convincing case that the protective role of streptokinase could have been confirmed ten years prior by aggregating the information from early studies. Early results were not truly contradictory; the studies were simply underpowered for the effect of interest – the ratio of the survival rates in the experiment and control arms. As you can see in this forest plot, the error bars in the early studies were huge, and overlapped 0, but many more studies showed a protective effect than not:

That the effect could have been uncovered earlier is obvious from the plot on the right, which shows a cumulative meta-analysis; what happens when we aggregate all the studies up to a certain study? It’s clear from this result that the evidence was already overwhelming by 1977.
A good meta analysis, such as this one, can:
- aggregate disparate information from different studies in an easily digestable from
- get precise estimates of effect sizes, allowing decisions to be centered around whether a treatment shows a clinically significant effect, as opposed to a simply statistically significant one
- identify points of agreement and disagreements between studies
- identify factors that moderate effect size through subgroup analysis and meta-regression
- guide the design of follow-up studies – for example, by identifying what is currently unclear in the extant data, or by furnishing an effect size estimate for power analysis
The mechanics of meta-analysis
The goal of a meta-analysis is to estimate an aggregate effect size. Now, different studies differ in substantive details – the target population, the randomization process, the practices of the institutions that perform the study – and so one cannot simply integrate data from different studies as though they’re coming from the same study.
Instead, one assumes that the true underlying effect – on survival, blood pressure, choice probability, etc. – differs from study to study. Even if each study had an infinite number of subjects, there would still be measurable discrepancies between the measured effect sizes between studies. These differences in effect sizes would then reflect substantive differences between the studies.
Under this random effects model, bigger studies count more, but smaller studies are still informative about the range of effects that one would measure in real life. The mean effect size is just as interesting as the range of effect sizes. An heterogeneity statistic, $I^2$, can help one judge whether studies generally agree or disagree, and whether this has substantive implications.
To identify the source of heterogeneity, one can use subgroup analysis and meta-regression. In subgroup analysis, you slice the data along dimensions of interest, e.g. smokers vs. non-smokers, or different treatment arms. In meta-regression, you instead regress a continuous variable against the outcome.
These approaches can be useful in identifying future directions of research, but they must be carefully interpreted, as they are correlational; even if each study was double-blinded, subgroup analysis and meta-regression will break randomization, and should not be interpreted causally.
Philosophical issues
The book goes into considerable detail into the manual computation of random effects models using a spreadsheet program, but in all honesty, you’re better off skipping these chapters, going straight into R and install.packages("meta"). I’m using this package for an ongoing meta-analysis I’m running; you can take a peek at what the code looks like here.
It’s in the discussion chapters, on philosophical issues and interpretation and gotchas that the book really shines. One chapter I really liked is on identifying publication bias. Studies can suffer from the file drawer problem: studies with negative results never get published because of a lack of incentive to do so.
There’s some techniques that have been developed to deal with publication bias. The underlying model is that large studies get published no matter their result, because they require so much resources and collaboration and are guaranteed a publication slot. However, smaller studies are much more prone to publication bias; who wants to publish a tiny study with negative results?
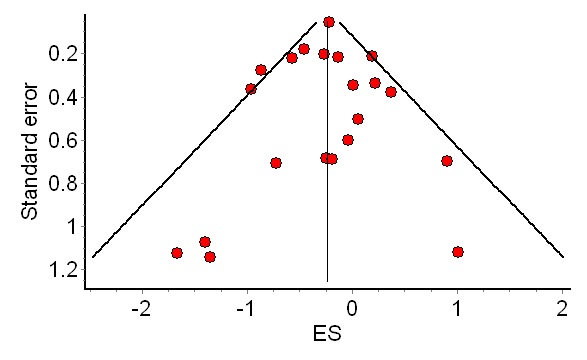
The funnel plot is a graphical tool to diagnose the presence of missing studies, which plots effect size on the x axis and study size on the y axis. Large studies are shown at the top, smaller studies at the bottom. Studies should be organized in an orderly funnel – more consistency for larger studies, and less so for the smaller studies. But if smaller studies are missing because of the file drawer problem, the funnel will be asymmetric; small studies with null results will be missing from the results.
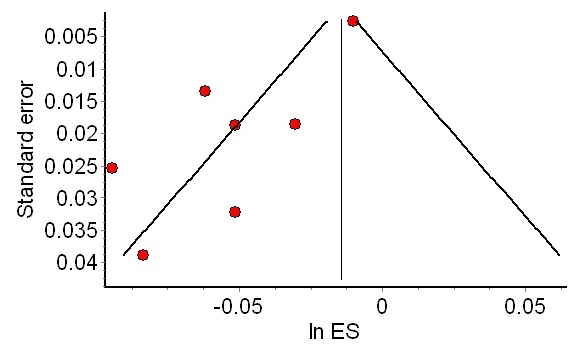
The authors a very nice, pragmatic method to estimate publication bias. The idea is to sort studies by descending size, and perform a cumulative meta-analysis; what happens if I analyze only the largest study? the two largest studies? the top 3? and so on. Mechanically, it’s the same analysis to the one that was applied to study when streptokinase could have been shown to be efficacious, but sorting by decreasing study size rather than date.
If the analysis suffers from publication bias, the effect size estimate should drift as one adds smaller and smaller studies to the meta-analysis. Rather than try and correct for the bias, one can simply highlight the sensitivity of the analysis to publication bias.
Closing thoughts
I highly recommend picking up this book as a first-line reference. I just wish it had a better discussion of dealing with missing data. I’m running a meta-analysis on glaucoma treatments with Dr. Masis these days and it seems like every other cell in my worksheet is NA. That’s unavoidable in a world where multiple researchers do independent studies without consulting each other; they’re going to measure slightly different things.
To deal with missing data in a principled way, we can use multiple imputation, or go full Bayesian. Each of these is its own large area of research, and I’ll follow up this post with my experiences with this.