I’ve used OpenBUGS for Gibbs sampling in the past, and while it’s a powerful piece of software, I hate OpenBUGS’ GUI, especially since it runs only on Windows. JAGS is an alternative Gibbs sampling program that uses an OpenBUGS-like syntax and runs on the command line. You can call JAGS in R through the rjags package and analyze the convergence of the MCMC chains through the coda package. It works beautifully, especially with RKWard, an alternative GUI for R that runs on Linux that is quite satisfying to use.
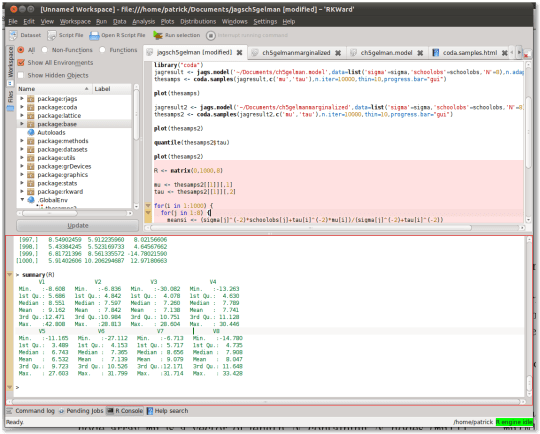
While its features do not rival that of Matlab — it doesn’t support debugging, for example –, it certainly eases you into the R language with features like syntax highlighting, inline help, and menus to perform common statistical analyses. Here’s a screenshot:
With RKWard and this Matlab to R cheat sheet, you can get started doing Gibbs sampling painlessly in minutes.
Installation
You can install JAGS and RKWard in Ubuntu at the terminal:
sudo apt-get install jags rkward
Then after opening RKWard you can install the “coda” and “rjags” packages through the Settings > Configure Packages menu, or by running the R command install.packages().
Simple example
Here’s a simple example taken from chapter 5 of Bayesian Data Analysis by Gelman et al. The problem is to determine whether different SAT training programs have a positive effect on SAT scores. We are given the summary statistics of the distribution of changes in SAT scores following training in 8 schools (mean and standard deviation), and they are quite noisy. Because of this noise, we might want to pool the scores from the different schools together to determine the overall effect of training, despite the fact that the schools used different programs. On the other hand, if we want to determine the effect in any individual school we can’t pool the results, and that means individual inference will be very noisy.
Gelman proposes that instead of either pooling everything together or doing no pooling at all, we should use a hierarchical model in which the mean training effect is taken from a distribution with mean 0, a school’s individual effect is taken from a distribution with mean equal to the mean training effect, and the observed scores are taken from distributions centered around individual effects. By tweaking the standard deviation (sigma) of the distribution of the school’s individual effects, we can obtain the equivalent of no pooling (sigma -> Infinity) or full pooling (sigma -> 0), or something in between.
Defining the model with JAGS syntax
The model is defined in JAGS syntax, which is similar to that of OpenBUGS, as:
model {
for(i in 1:N) {
schoolmean[i] ~ dnorm(mu,itau)
thes[i] <- 1/pow(sigma[i],2)
schoolobs[i] ~ dnorm(schoolmean[i],thes[i])
}
mu ~ dnorm(0,alpha)
alpha <- .01
itau ~ dgamma(1e-3,pow(15,2)*1e-3)
tau <- pow(1/itau,1/2)
}
This model can be saved as ~/Documents/ch5gelman.model.
Running the model in R
We can write the data (Table 5.2 in Gelman) in R syntax as:
sigma <- c(15,10,16,11, 9,11,10,18) schoolobs <- c(28,8, -3, 7,-1, 1,18,12)
To run the model, we include rjags and coda through the library command, and write:
library("rjags")
library("coda")
jagresult <- jags.model('~/Documents/ch5gelman.model',data=list('sigma'=sigma,'schoolobs'=schoolobs,'N'=8),n.adapt = 1000)
thesamps <- coda.samples(jagresult,c('mu','tau'),n.iter=10000,thin=10,progress.bar="gui")
You have just defined a hierarchical Bayesian and Gibbs sampled from it! Pretty easy, right? jags.model compiles the model while coda.samples samples from it. Did the Gibbs sampler converge though? You can visualize the chain using plot(thesamps):
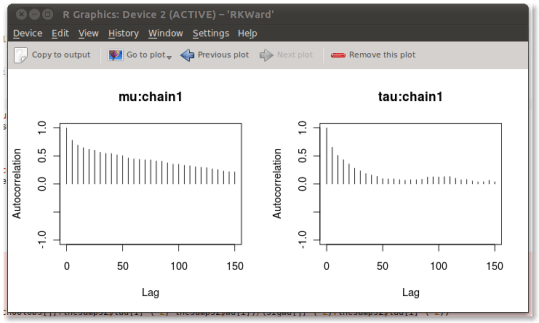
That looks terrible. Clearly this chain has not converged; mu looks terrible, in particular. You can do a smörgåsbord of convergence diagnostics through a wizard-like interface with codamenu(). This allows you, for example, to plot the autocorrelation of the chain:
Clearly, this won’t do.
A collapsed Gibbs sampler
Our chain exhibits poor convergence because mu and schoolmean are heavily correlated. This issue can be avoided by simply integrating out schoolmean, thus creating a collapsed Gibbs sampler. As this is a very simple model, this is straightforward:
model {
for(i in 1:N) {
thes[i] <- 1/(pow(sigma[i],2) + 1/itau)
schoolobs[i] ~ dnorm(mu,thes[i])
}
mu ~ dnorm(0,alpha)
alpha <- 1e-4
precisiontau <- .001
meantau <- 15
itau ~ dgamma(precisiontau, precisiontau*pow(meantau,2))
tau <- 1/sqrt(itau)
}
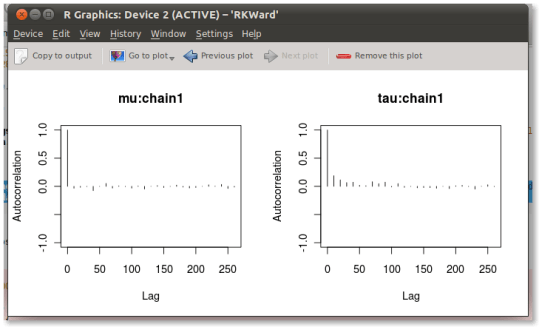
Running this new model, you can see that the autocorrelations are much better now:
Unfortunately, since we’ve removed schoolmean from the model through integration, we can’t get an estimate for the school means. You can, however, sample from the distribution of schoolmean in R using the conditional distribution of schoolmean given everything else (equation 5.17):
mu <- thesamps[[1]][,1]
tau <- thesamps[[1]][,2]
for(i in 1:1000) {
for(j in 1:8) {
meansi <- (sigma[j]^(-2)*schoolobs[j]+tau[i]^(-2)*mu[i])/(sigma[j]^(-2)+tau[i]^(-2))
stdsi <- (sigma[j]^(-2)+tau[i]^(-2))^(-1/2)
R[i,j] <- rnorm(n = 1,mean=meansi,sd=stdsi)
}
}
You can then use functions like quantile and summary in R to obtain summary statistics for R. For me, this results in:
> summary(R) V1 V2 V3 V4 Min. :-8.608 Min. :-6.836 Min. :-30.082 Min. :-13.263 1st Qu.: 5.686 1st Qu.: 4.842 1st Qu.: 4.078 1st Qu.: 4.630 Median : 8.551 Median : 7.597 Median : 7.260 Median : 7.789 Mean : 9.162 Mean : 7.842 Mean : 7.138 Mean : 7.741 3rd Qu.:12.471 3rd Qu.:10.984 3rd Qu.: 10.751 3rd Qu.: 11.128 Max. :42.808 Max. :28.813 Max. : 28.604 Max. : 30.446 V5 V6 V7 V8 Min. :-11.165 Min. :-27.112 Min. :-6.713 Min. :-14.780 1st Qu.: 3.489 1st Qu.: 4.153 1st Qu.: 5.717 1st Qu.: 4.735 Median : 6.743 Median : 7.365 Median : 8.656 Median : 7.908 Mean : 6.532 Mean : 7.139 Mean : 9.079 Mean : 8.047 3rd Qu.: 9.723 3rd Qu.: 10.526 3rd Qu.:12.171 3rd Qu.: 11.648 Max. : 27.603 Max. : 31.799 Max. :31.714 Max. : 33.428
You can see that compared to Table 5.3 in Gelman, the means are shrinked more towards the population mean as I have a proper prior on tau rather than a uniform one as in his example.

5 responses to “Gibbs sampling made easy – JAGS, RKWard, CODA”
Thanks for your great work! I wonder whether I can use the same way of collapsed Gibbs Sampling to fit the LDA model? Thank you in advance!
[…] exactly onto the per-school mean SAT example from Gelman et al., chapter 5. I had written up about getting this model going in R about 4 years ago, and it still runs. Here’s what the shrunken estimates look […]
Hi, thanks for this post. The rjags help says thatt function coda.samples updates the model, and coerces the output to a single mcmc.list object. I would like to know how to update it again if you haven’t reach convergence without loosing the previous iterations.
Thanks for your answer!!!!!
I don’t know, I guess you could simply concatenate the mcmc.list objects you get on every update.
Excelent post. I’m really astonished bugs doesn’t have natural gui’s under linux neither a port into CRAN (at this time) although I have installed OpenBUGS, but run this into shell it is a… ‘different experience’.