Decoding neuronal activity is a powerful technique to study how information is encoded in a population and how it might be extracted by other brains areas. Hippocampal place cells are a prime example of a system that can be studied fruitfully from a decoding persepective.
In a typical place cell decoding experiment, a population of neurons is recorded while an animal (typically a rat) is moving in a field. The movements of the animal are recorded using a camera. The decoder is then trained using a dataset containing fully observed data (both spikes and positions).
The decoder is then tested on another dataset by predicting the location of the animal based solely on the recorded neural data. Typically, the decoder is constrained to use only causal information to increase the relevance of the analysis (since neurons can’t read spikes that haven’t occurred yet). So if is a vector representing the position of the animal at time t, the goal of the decoder is estimate
, where
is the response of the j’th neuron in time bin m.
Typically the problem is solved by continuous state-space methods involving recursive filtering like the Kalman filter, the extended Kalman filter (EKF), unscented Kalman filter (UKF), or more generally sequential Monte Carlo methods (particle filters). I’m interested in learning particle filter methods, but I figured I should warm up with a deterministic decoding algorithm.
Hence I decided to implement the algorithm proposed by Emery Brown et al. (1998; A statistical paradigm for neural spike train decoding applied to position prediction from ensemble firing patterns of rat hippocampal place cells, J Neurosci). I decided to ignore the theta phase component of their model to focus on position encoding and decoding.
To decode the activity of a cell ensemble, you must first specify an encoding model, that is, a model that specifies how the firing pattern of neurons is related to the stimulus. Brown et al. make the straighforward assumption that the number of spikes in a bin for the j’th neuron is related to the proximity of the animal to the cell’s Gaussian-shaped place field. The output of the Gaussian gives the cell’s mean firing rate and its measured response is derived from a Poisson process:
Here is a 2d vector specifying the preferred position of the cell,
is a 2d vector specifying the extent of the RF in the x and y directions, and
if the offset of the cell. Keen observers will notice that this is actually a Quadratic-Exponential-Poisson model, that it is formally equivalent to the model underlying Bayesian spike-triggered covariance (BSTC), and that its parameters can be fit through Maximum Likelihood in a standard exponential/Poisson GLM.
For the purposes of the demonstration, however, I will simply draw random parameters for the place cells and use the true values in the decoding process.
Another component that must be specified to do decoding is the prior probability distribution of the parameters to be decoded. A standard default assumption is that the parameters are generated by a low-order autoregressive process (typically a first-order Markov chain). In Brown et al. the authors specify that the change in position from bin to bin is generated by iid. Gaussians:
Here W is the covariance matrix of the change in position from frame to frame. Again, such data is easy to simulate; a simple call to randn, then cumsum will do the trick. Thus, I simulated a path through an open 2d environment and the response of 25 place cells like so:
%start by simulating a path in a 2d array ntimes = 1000; path = randn(ntimes,2)*.03; %path = conv2(path,ones(1000,1)/1000,'same'); %path = path/std(path(:)); path = cumsum(path); ncells = 25; %% cellparams = zeros(ncells,5); %centers cellparams(:,1:2) = randn(ncells,2); %sizes cellparams(:,3:4) = abs(randn(ncells,2)); cellparams(:,5) = randn(ncells,1)*.5; %% R = zeros(ntimes,ncells); for ii = 1:size(cellparams,1) x0 = cellparams(ii,1); y0 = cellparams(ii,2); sigmax = cellparams(ii,3); sigmay = cellparams(ii,4); r = exp(-(path(:,1)-x0).^2/2/sigmax^2-(path(:,2)-y0).^2/2/sigmay^2 + cellparams(ii,5)); R(:,ii) = poissrnd(r); end
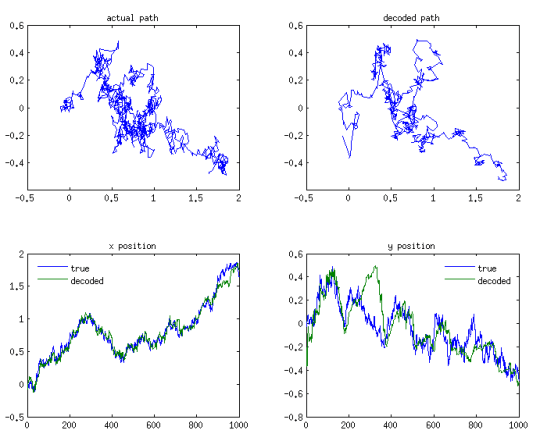
The simulated path is shown at the top left of the post, while the resulting spike trains are shown below:
Given this setup, the position of the rat can be decoded using these recursive equations (eqs. 6 and 7 in the original) which are straightforward applications of Bayes’ theorem:
Here K is an irrelevant constant (somehow WordPress’ LaTeX parser doesn’t handle \propto). What’s important to notice here is that the transition probability is a standard Gaussian distribution, while the posterior distribution
is a non-standard distribution. That means that we can’t use the standard equations of the Kalman filtering framework to perform the necessary recursions, since this requires a Gaussian posterior and a Gaussian transition probability.
A simple remedy is to approximate with a Gaussian distribution. This can be done most simply by taking
, where
is the Maximum a Posterior (MAP) estimate of the position, and
is the Hessian of the log-posterior at the MAP estimate. This is called the Laplace approximation.
Such is the basis of the iterated extended Kalman filter (IEKF). Iterated in this context refers to Newton iterations; for one Newton iteration you get something similar to an extended Kalman filter (I think; I’m kind of confused by all the different variants of the EKF). A relevant reference is this thesis by Bjarke Mirner Klein on State space models for exponential family data. He has an R library available that performs IEKF (although I think it only applies to a strict GLM without quadratic terms, so it wouldn’t work here).
Given this approximation, recursive filtering is straightforward. If , then the second equation is a convolution of a Gaussian with another zero-mean Gaussian, and hence:
All this means is that going forward in time by one step spreads the posterior probability of the position by an amount proportional to the expected size of the changes in speed. Then to determine , we perform an optimization; it looks like the log-likelihood of a GLM with a quadratic penalty. From this, we extract the MAP estimate and the Hessian. I did the optimization through minFunc because I was too lazy to write a damped Newton solver. to verify the gradient and Hessian.
By applying the steps recursively, we obtain a causal estimate of the position of the rat as a function of time, as illustrated above. The whole thing is surprisingly straightforward, with the main loop taking only 10 lines of code or so. The trickiest bit was getting the correct gradients and Hessian while sleep-deprived; I used the derivest suite to verify my results. As you can see, the resulting estimates (shown at the top of the post) are quite accurate despite the small number of cells. Next, I’ll try to get similar results with a particle filter.
Here is the decoder I wrote:
function [xs,Ws] = brownDecoder(Y,params,W)
%R is the response matrix (Ntimesteps x ncells)
%params are the parameters of the cells (ncells x 5, corresponding to
%x, y, sigmax, sigmay, and offset)
%W is the covariance matrix from p(x_k|x_k-1) = N(x_k-1,W^-1)
xhat = [0;0];
What = eye(2)*5;
xs = zeros(size(Y,1),2);
Ws = zeros(size(Y,1),2,2);
mfopts.Method = 'newton';
mfopts.Display = 'off';
mfopts.TolX = 1e-4;
mfopts.TolFun = 1e-4;
for ii = 1:size(Y,1)
%Computing p(x(t_k)|spike in (0,t_k-1)) is easy, it's simply
%N(xhat, What + W)
What = What + W;
%Next, figure out p(x_k|y_1...y_k) \approx N(xhat, What)
%through optimization
y = Y(ii,:)';
Wihat = inv(What);
xhat = minFunc(@(x) computeStatePosterior(x,y,params,xhat,Wihat),xhat,mfopts);
[~,~,Wihat] = computeStatePosterior(xhat,y,params,xhat,Wihat);
What = safeInverse(Wihat);
xs(ii,:) = xhat';
Ws(ii,:,:) = What;
if mod(ii,100) == 0
fprintf('Iteration %d\n',ii);
end
end
end
function [Hi] = safeInverse(H)
[~,p] = chol(H);
if p > 0
H = H + eye(size(H,1)) * max(0,10 - min(real(eig(H))));
end
Hi = inv(H);
end
function [E,g,H] = computeStatePosterior(x,y,params,xhat,Wihat)
%Predicted rate for each neuron given Gaussian RF models
loglambdas = -1/2./params(:,3).^2.*(x(1)-params(:,1)).^2 + ...
-1/2./params(:,4).^2.*(x(2)-params(:,2)).^2 + params(:,5);
lambdas = exp(loglambdas);
%Compute negative log-posterior
%First part is due to likelihood of data given position, second part is
%the prior prob. of positions
E = sum(-y.*loglambdas + lambdas) + 1/2*(x-xhat)'*Wihat*(x-xhat);
if nargout > 1
%Compute gradient of error (first part: likelihood)
dloglambdas = [-1./params(:,3).^2.*(x(1)-params(:,1)),-1./params(:,4).^2.*(x(2)-params(:,2))];
g1 = zeros(2,1);
g1(1) = sum(-y.*dloglambdas(:,1) + lambdas.*dloglambdas(:,1));
g1(2) = sum(-y.*dloglambdas(:,2) + lambdas.*dloglambdas(:,2));
%Second part: prior
g2 = Wihat*(x-xhat);
g = g1+g2;
end
if nargout > 2
%Compute Hessian of error, first part
H1(1,1) = sum(-y.*(-1./params(:,3).^2) + lambdas.*dloglambdas(:,1).^2 + lambdas.*(-1./params(:,3).^2));
H1(1,2) = sum( lambdas.*dloglambdas(:,1).*dloglambdas(:,2));
H1(2,1) = H1(1,2);
H1(2,2) = sum(-y.*(-1./params(:,4).^2) + lambdas.*dloglambdas(:,2).^2 + lambdas.*(-1./params(:,4).^2));
H2 = Wihat;
H = H1 + H2;
end
end
Called like so:
W = eye(2)*.03^2; [xs,Ws] = brownDecoder(R,cellparams,W);